ILMSImage Attribute Calculation
Contents |
 ILMSImage Attribute Calculation
ILMSImage Attribute Calculation
Introduction
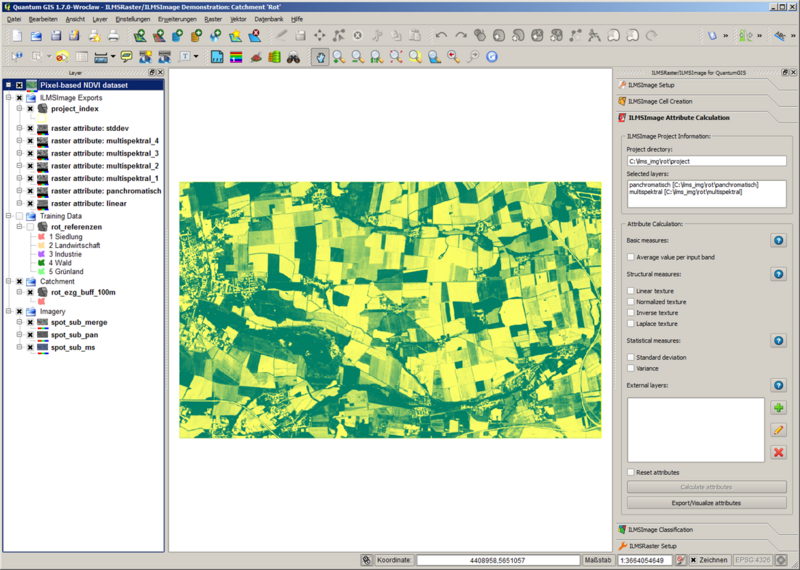
ILMSImage Attribute Calculation is part of the ILMSImage plug-in for QuantumGIS and is used for the derivation and calculation of cell-based attributes and for the integration of additional data layers.
Similar to various ILMSImage panels it consisits of two components, ILMSImage Project Information in the upper section and the actual tools in the lower section.
Attribute Calculation
Background
...
Overview
The panel allows the following functions for the user:
- Calculation of default cell-based attributes: In addition to features of cell geometry basic attributes, different structural features and statistical measures belong to the implemented parameters. After the calculation those are summarized in form of an attribute table.
- Adding additional data layers: The corresponding data layers are selected by the user, an identifier is assigned and added to the existing data layers. If necessary, the data type is automatically adapted and the selected additional data are cut in terms of space.
- Visualization of selected attributes: For this purpose the attributes which are summarized in a database table are transformed into raster data and prepared in QuantumGIS for visualization.
Calculation of Cell-Based Attributes
In order to derive specific attributes for the samples from the link of cell geometry and selected input data, the relevant parameters have to be activated by clicking on the corresponding check box. The default attributes are basic values, structural features, statistical parameters and the implicitly calculated features of cell geometry.
| Attribute type | Attribute | GUI-Label | Internal designation | Description |
|---|---|---|---|---|
| Basis | Average value per band | average value per input band | <layer_id>[_<numerator>] |
For every input band an average value is calculated from all image pixels summarized in one cell. The internal designation corresponds to the identifier which is selected for the corresponding band (in the example: panchromatic), in case of mult-channel input data completed by one channel number (in the example: multi-spectral_1, multi-spectral_2, etc.).
|
| Structure | Linear texture | linear texture | linear |
The activation of this measure creates the 1st degree texture in a 3x3 kernel. The algorithm calculates the texture for every cell individually and ignores all image pixels which are outside of the current cell. Like all structural features also the linear texture uses the first channel which is displayed in the project information as a basis for calculation. |
| Normalized texture | normalized texture | normalized |
The normalized textures uses the same algorithm as the linear texture but it normalizes the result additionally with the average value of all image pixels of the current cell. Like all structural features also the normalized texture uses the first channel which is displayed in the project information as a basis for calculation. | |
| Inverse texture | inverse texture | inverse |
By activating the inverse texture the Inverse Difference Moment (Haralick et al. (1973), S. 619) is calculated on pixel level. Inverse texture uses the same algorithm as the linear and normalized texture but it finally calculates the inverse value of the square differences. Like all structural features also the inverse texture uses the first channel which is displayed in the project information as a basis for calculation. | |
| Laplace texture | Laplace texture | laplace |
This structural feature uses the modified Laplace kernel for contrast enhancement for individual cells. The parameter enhances the result with a generic kernel which reacts to local contrast maxima with minimal spatial distance (two to five pixels). As a result, isolated lines are reduced and regions are filled with spatially dense contrasts. | |
| Statistics | Standard deviation | standard deviation | stddev |
By activating this option the standard deviation of all image pixels summarized in one cell of the first channel in the project information is calculated. |
| Variance | variance | variance |
By activating this option the variance of all image pixels summarized in one cell of the first channel in the project information is calculated. | |
| Cell geometry | cell size | without GUI link | size |
The size of one cell in image pixels of the most highly resolved input data channel. |
| Cell perimeter | perimeter |
The perimeter of one cell in image pixels of the most highly resolved input data channel. | ||
| Cell length | length |
The length of a cell in image pixels of the most highly resolved input data channel. | ||
| Cell width | width |
The width of a cell in image pixels of the most highly resolved input data channel. |
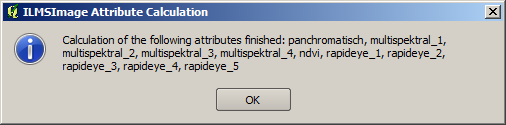
After the selected attributes have been calculated, they are summarized in a table and are available for additional analysis using ILMSImage. The user receives a note about the end of the process and a summary of the calculated attributes.
Export and Visualization of Cell-based Attributes
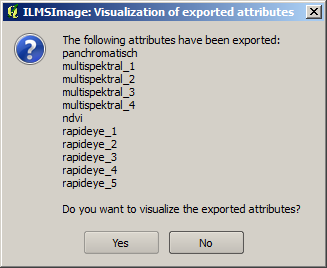
For visual control the plug-in makes it possible to export selected attributes as raster layers after they have been calculated and to show them subsequently in the current QuantumGIS-Project. Consequently, the required attributes are selected and the process is started by clicking on Export/visualize attributes. Then ILMSImage creates raster datasets of the attributes and stores them in the exports directory of the current ILMSImage project. The completion of the process is indicated to the user in a message window.
The window also offers the possibility of visualizing all exported attributes automatically. For this purpose the raster datasets are loaded into the current QuantumGIS project, relocated to the key group for ILMSImage Exports and their visualization is adapted by using a Standard Deviation Stretch. The result is represented in a map view like this:
If the corresponding prompt is denied, the exported attributes can be added to the current QuantumGIS project individually and separately by the user. In this case, the visualization has to be modified by the user as well.
Working with External Data Layers
Adding and Editing Additional Data Layers
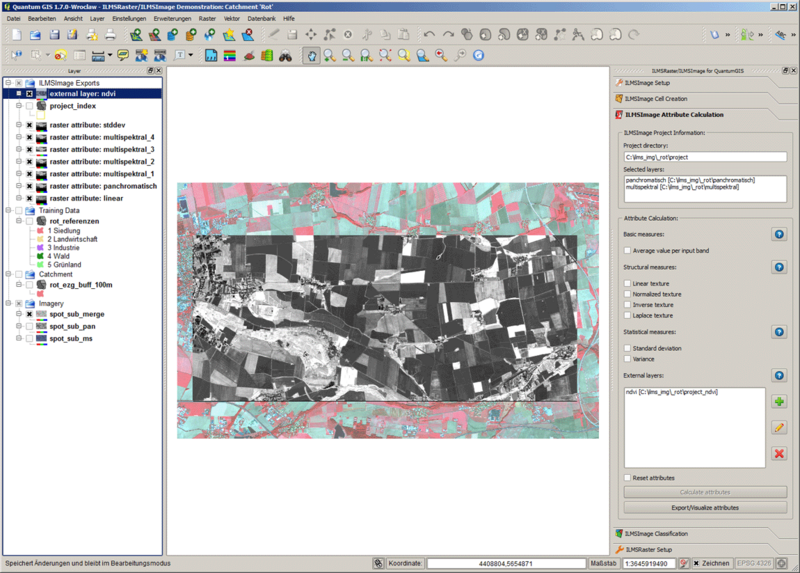
In addition to the mentioned standard attributes, the external data layers can be integrated into the process of attribute calculation. For the integration basically every available dataset in raster form is suitable. In the integration within every layer and for every cell a mean value is calculated and finally added to the attribute table. This means that additional attributes correspond to the above-described basic attribute - with regard to the calculation ordinance – the only difference being that additional data layers do not influence cell creation since they are already completed. This function is therefore suitable for integrating additional multispectral levels (or combinations of these) which are required as attributes for additional analysis but not for a derivation of analysis areas, i.e. cells, into the work process. A simple example is the Normalized Vegetation Difference Index, NDVI which is a measure for vegetation activity. It can be calculated using two channels which are in the visual field and in the close infrared field of the electromagnetic spectrum. As additional data layer a pixel-based NDVI picture is available. The following image shows areas of high vegetation activity in green colors and areas of low activity in yellow colors.
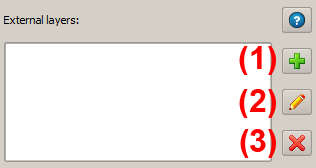
This dataset can now be defined as an external data layer - for this purpose it does not have to be loaded in QuantumGIS; it is sufficient to know the corresponding path. For the definition the corresponding field of the panel is used. The tool allows to define new external layers (1), edit existing layers (2) and delete external data layers from the project (3).
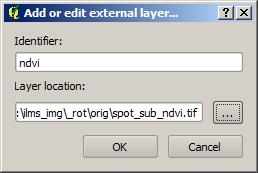
In order to create a new external layer, a dialog box appears after clicking on the corresponding icon. It makes it possible to select the corresponding data source in the file system and assign an identifier. It should be noted that a unique identifier is selected which is not already used for other data layers or for one of the default attributes (see table above). For the example dataset of NDVI it looks like this:
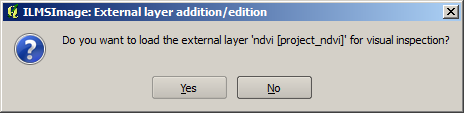
After the affirmation the dataset is copied to the project directory to allow ILMSImage exclusive access to the source files for cell creation in an analogous manner. If necessary, it is then adjusted to the test area. In order to allow for visual control, ILMSImage offers to load the results of these processes into the map view:
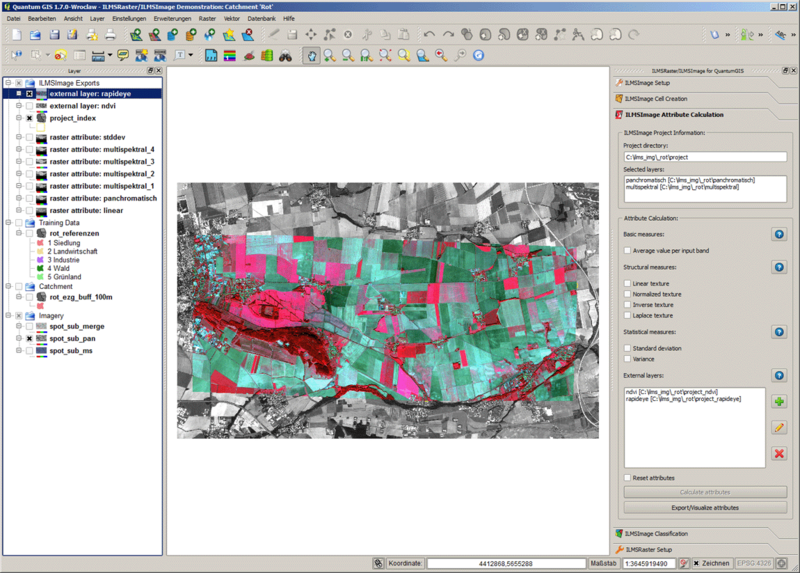
After affirming the question, the external dataset is visualized using a Standard Deviation Stretch; the range of colors for original data is not applied. Since this representation is only used for visual control, the corresponding dataset is deleted before carrying out an attribute calculation. Moreover, the generated additional level appears in in the corresponding list of the tool panel. In the following image an area of high vegetation activity is shown in light colors, areas of low activity are represented in dark colors:
A dataset which has already been established can be edited by clicking on the corresponding symbol. It is possible to change the identifier and modify the path of the source file.
Automatic Scaling of the Data Type of External Data Levels
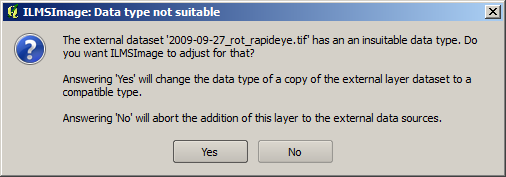
ILMSImage only allows input data which are available in the byte domain (8bit, gray level between 0 and 255). If a non-compatible layer is chosen during the definition of an external data level, ILMSImage asks if this layer is supposed to be scaled automatically on the acceptable domain. If, for example, a 16bit part of a RapidEye scene is supposed to be integrated, the following message appears:
When the request has been confirmed, the dataset is scaled on the 8bit domain and is loaded - optionally - into the map view for visual control.
Attribute Calculation Using External Data Layers
The defined external datasets are available for attribute calculation like the original source data did and are therefore treated like equivalent layers. This holds true for the actual calculation and the visualization of the calculated layers.
Deleting External Data Layers
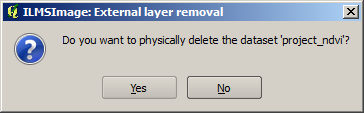
In order to remove an external data layer from the project, it is selected from the list of all external layers and the appropriate symbol in the panel is clicked. ILMSImage then asks whether the related dataset should be deleted permanently, i.e. removed from the file system. This request refers only to a generated rush print of the external layer which was generated earlier in the project directory. The files which were originally used for the definition are not threatened with deletion.
Following Steps
After cell creation a typical ILMSImage process chain with derivation and calculation of cell attributes continues by using the panel ILMSImage Classification.